Bacterial Translocation and Changes in the Intestinal Microbiome in Mouse Models of Liver Disease
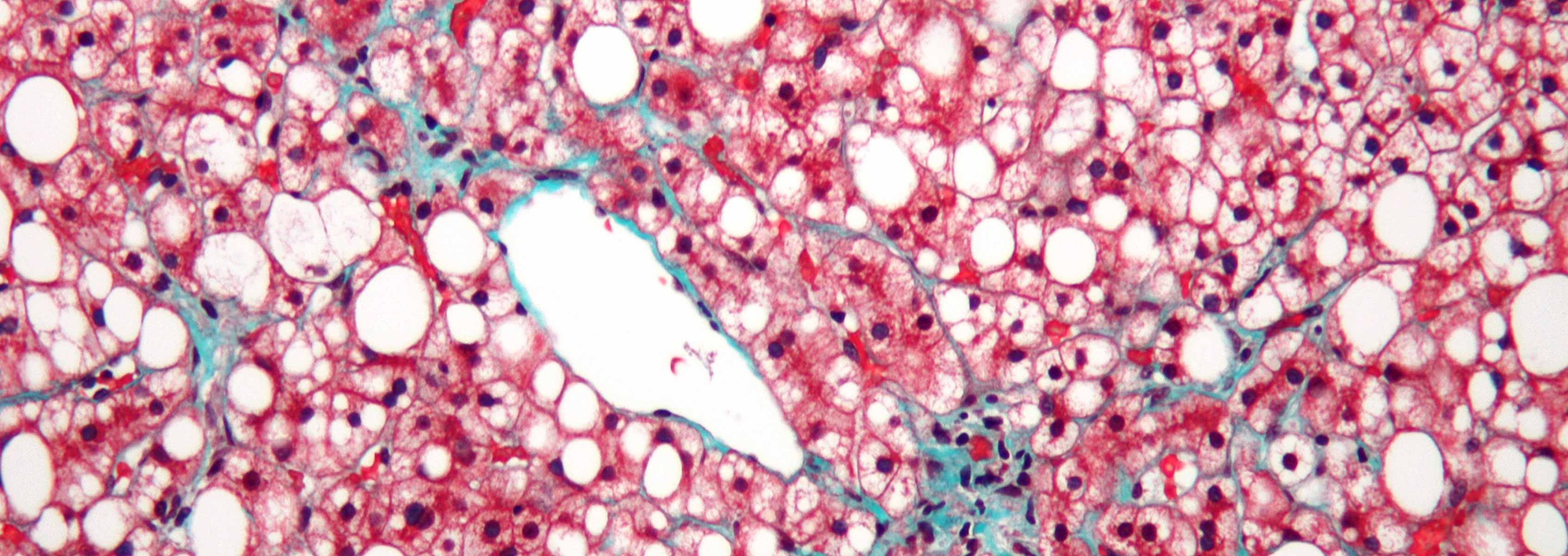
According to the Centers for Disease Control and Prevention, mortality associated with chronic liver disease ranks as the 12th most common cause of death in the U.S., but may be as high as 8th if obesity-related fatty liver disease, viral hepatitis, and liver cancer are included in the equation. Patients with later stages of chronic liver disease (e.g. fibrosis, cirrhosis) have bacterial overgrowth in the small intestine, and translocation of gut bacteria and their products from the lumen of the intestine to mesenteric lymph nodes, blood, and extraintestinal organs. Animal models that mimic the disease found in humans are very powerful tools for understanding disease progression and discovering potentially novel treatment interventions for humans. To understand the changes in the intestinal microbiome and bacterial translocation during early stages of liver injury, three mouse models of liver disease were used in two separate studies.
In the first study, published in Hepatology, we used a mouse model of continuous intragastric feeding of alcohol with mice fed a non-alcoholic diet of similar caloric intake as controls. The aims of this study were to investigate bacterial translocation, changes in the intestinal microbiome, and expression of intestinal antimicrobial proteins in alcoholic liver disease. Intestinal bacterial overgrowth was observed in the intestinal tract of mice fed alcohol for 3 weeks compared to control mice by conventional bacterial culturing techniques. Since most gut bacteria cannot be cultured, we used high-throughput 16S rRNA gene sequencing to assess the qualitative changes in the intestinal microbiome during alcohol feeding. The results of 16S rRNA gene sequencing revealed an increase in the relative abundance of the Gram-negative bacterial order Verrucomicrobia and Bacteriodetes, with a reduction in the Gram-positive order Firmicutes. Bacterial translocation, noted by increased levels of bacteria and endotoxin in blood, was elevated in alcohol fed mice compared to controls. We found that alcohol feeding resulted in decreased expression of bactericidal c-type lectins Reg3b and Reg3g. Treatment with prebiotic fructo-oligosaccharides resulted in restored Reg3g protein expression, reduced bacterial overgrowth, and lessened liver disease.
In the second study, published in J. of Hepatology, intestinal permeability, bacterial translocation, and the intestinal microbiome were studied in two mouse models of liver disease; cholestasis and toxic liver injury. Microbiome data was also compared between four different mouse models of liver injury. Ligation of the common bile duct (BDL), modeling cholestatic liver injury, resulted in a rapid onset of bacterial overgrowth; however, it took 24 injections of carbon tetrachloride (CCl4), modeling toxic liver injury, to observe bacterial overgrowth. Though there were differences in the timing of bacterial overgrowth between these two mouse models, an increase in intestinal permeability and bacterial translocation was observed in both models in just one day following treatment. These increases were accompanied by disruption of tight junctions and a decrease in expression of the tight junction protein occludin. Analysis of the intestinal microbiome, targeting the 16S rRNA gene, revealed minor changes following BDL, but major changes following CCl4. Specifically, there was an increase in the relative abundance of Firmicutes and Actinobacteria in treated compared to untreated mice. Shannon and Simpson indexes indicated a decrease in bacterial diversity in the CCl4-treated group. Comparison of microbiome data from the alcohol model, BDL, CCl4, and an obesity model noted that there was no common liver disease microbiome, suggesting that the intestinal microbiome differs by etiology of liver disease.
Funding
Financial support for these projects was in part through NIH grants P50AA11999, K08 DK081830, R01 AA020703, R01 DK072237, the UCSD Digestive Diseases Research Development Center (DK080506), and the J. Craig Venter Institute.